RNA Sequençing data analysis
The intricate mechanisms of gene regulation
Below we present some of the most common analyses we perform on RNA-seq data. The explorative, differential expression and pathway analyses largely apply to other high-throughput expression data as well, such as expression microarray or proteomic data.
Exploratory gene expression analysis
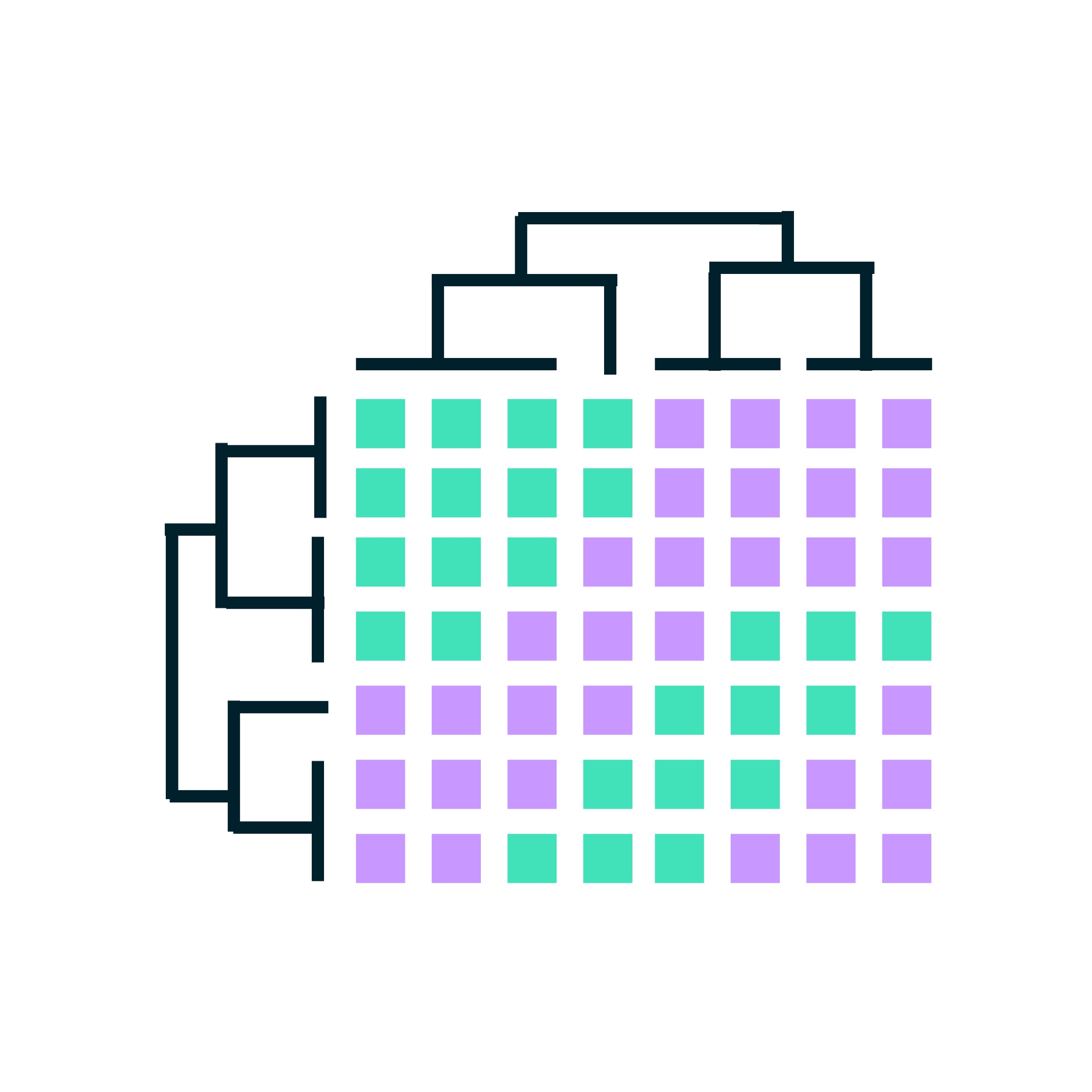
Every RNA-seq expression study incorporates an exploratory analysis. After the raw sequencing reads of an RNA-seq experiment have been quality controlled and gene counts derived, the data set is visualized using Principal Component Analysis (PCA) and expression heatmaps to unveil general patterns. These visualizations help us answer questions such as:
- Do the biological replicates resemble each other with regards to their expression profiles?
- Do distinct sample groups (e.g., different tissues, treatments or time points) form separate clusters?
- Are there outlier samples?
Differential expression analysis
Differential expression analysis is a statistical comparison of two (or more) sample groups. It results in differential expression statistics for each detected transcript, such as the fold change and statistical significance. These statistics are typically visualized using a volcano plot. The genes which are found to be up- or down-regulated can be further visualized as heatmaps or boxplots, for instance.
As a statistical analysis, this phase of an expression study benefits from the statistical power brought by biological replicates. Three biological replicates per condition is a common “rule-of-thumb” minimum, but it only allows for reliable detection of genes with relatively large expression differences. With a careful experimental design and sufficient sample size, subtler differences can be detected and confounding factors controlled for.

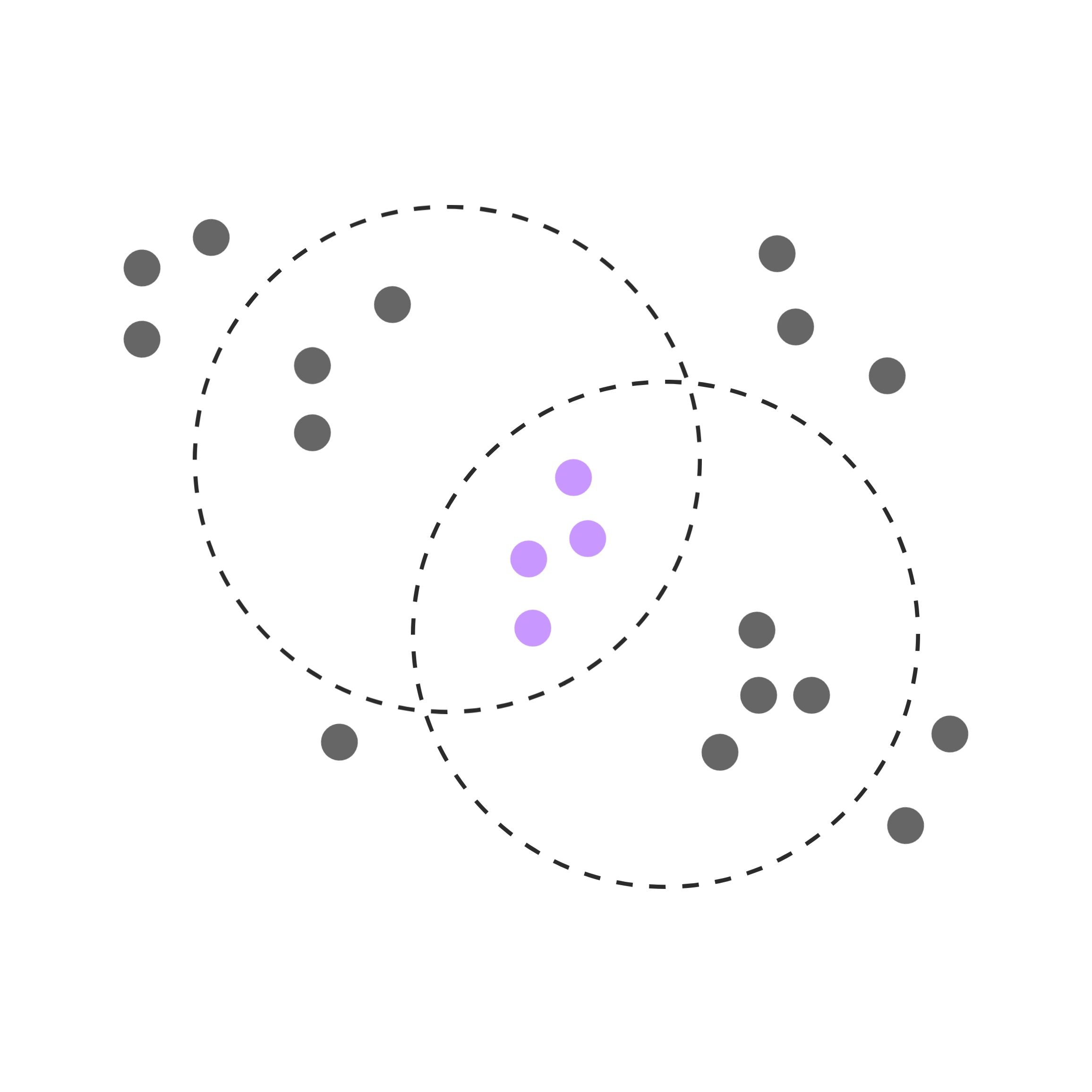
Pathway analysis
Pathway analysis puts genes from a differential expression analysis into broader biological context. Simple pathway analyses compare the up- and down-regulated genes statistically to predetermined gene lists. These lists are annotated to biologically meaningful terms, such as a biological process, signaling pathway or a specific disease.
Such analyses may rely either on over-representation analysis or gene set enrichment analysis, which both result in a list of enriched gene sets with relevant statistics and annotations.
More mechanistic pathway analyses rely on experimentally validated interactions between genes. They enable identifying not just which pathways are represented by the differentially expressed genes, but also shed light on whether the pathways are activated or inhibited, and by which genes.
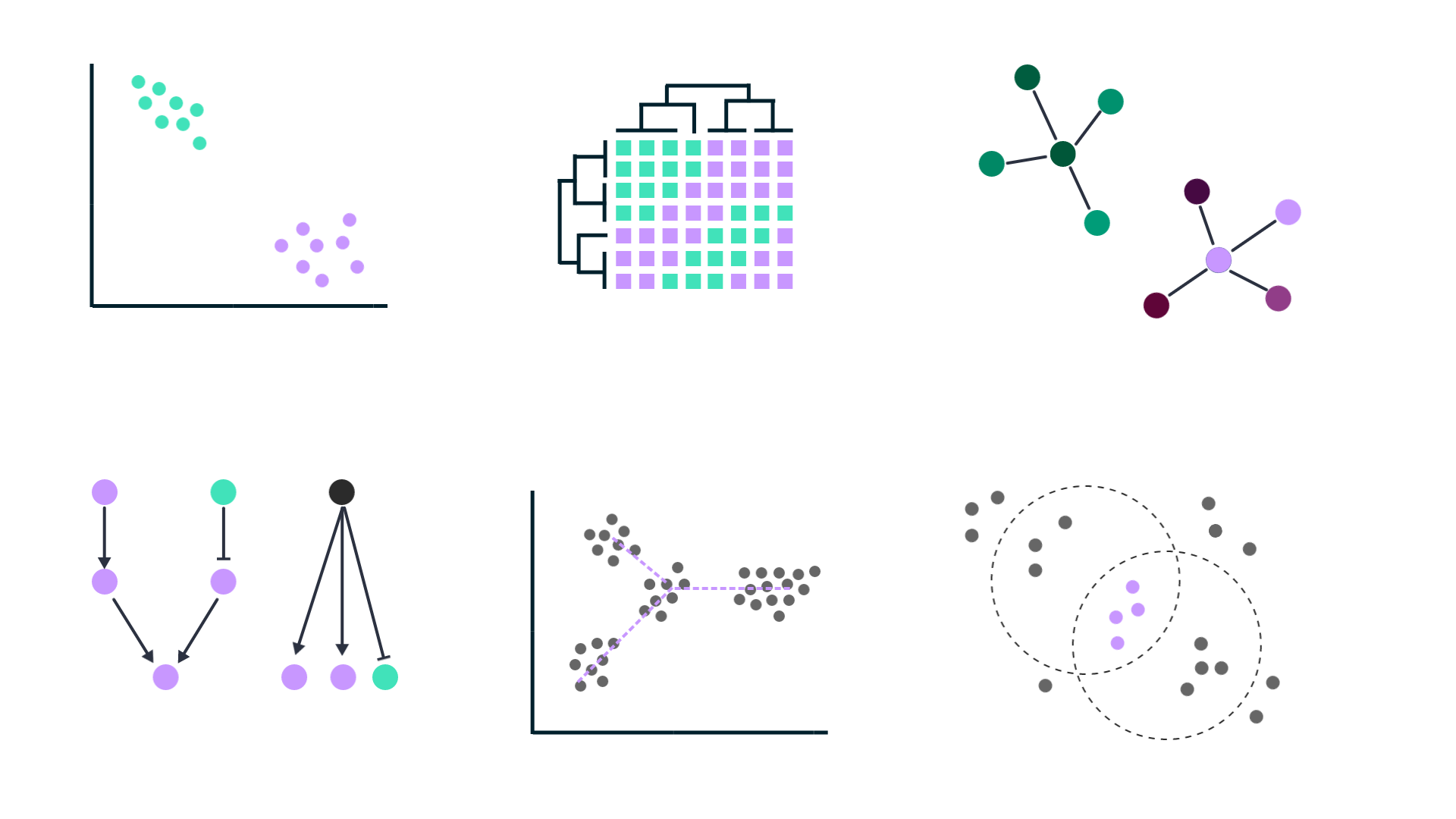
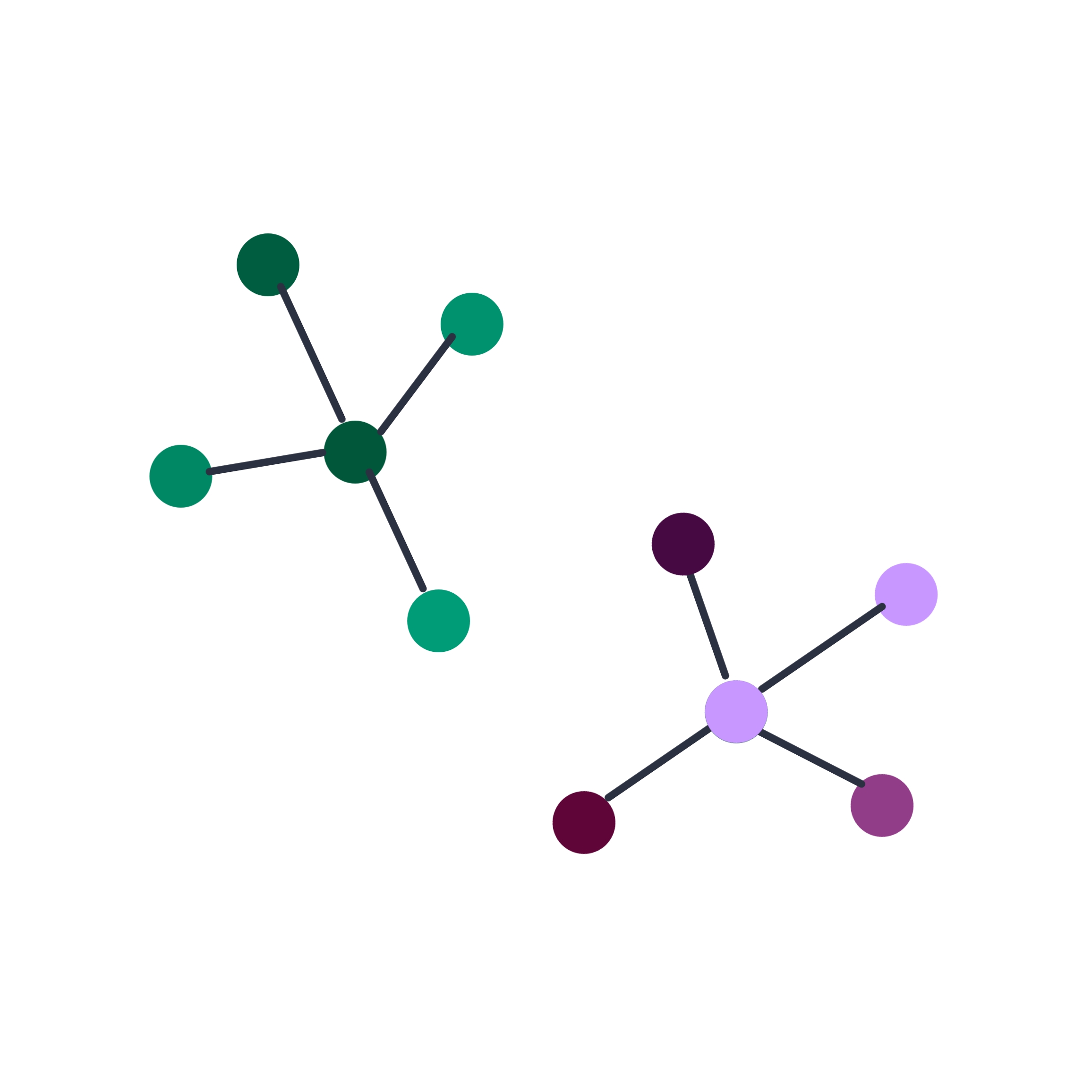
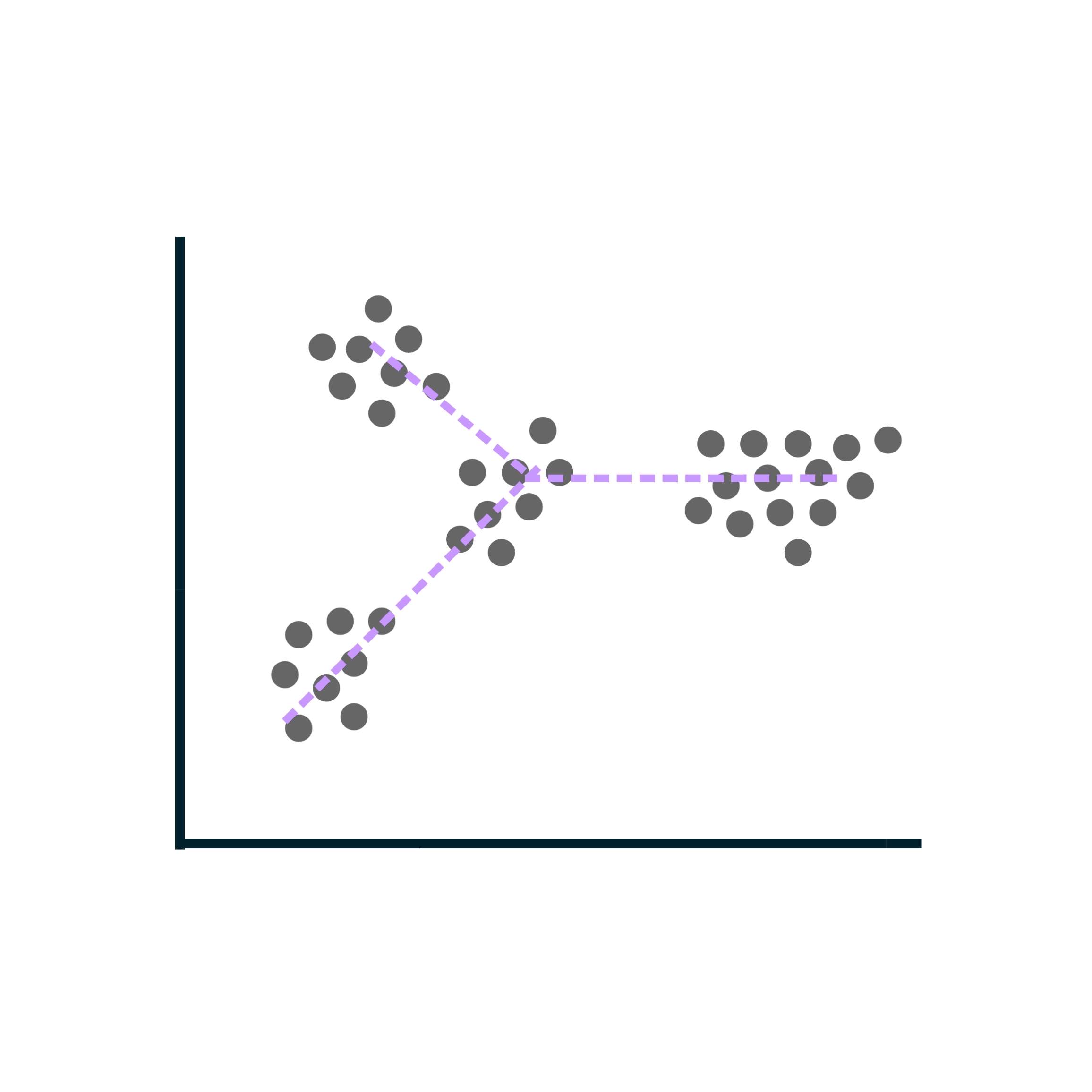
Single-cell expression analysis
Single-cell RNA-sequencing (scRNA-seq) experiments allow for cataloguing cell types and uncovering differentiation trajectories at a scale and resolution unmatched by bulk RNA sequencing.
Used particularly to study the composition and development of complex tissues, scRNA-seq data sets typically comprise thousands of individual cells. Most approaches used to analyze bulk RNA-seq data can be tailored for single-cell RNA-seq data as well.
STEP1 - Contact us
We have a meeting, phone call or visioconference to discuss your project
STEP2 - Estimate and timeline
Our team compiles a project proposal for your project
STEP3 - Accept or reject
The proposal is up for your consideration
STEP4 - Let Magic Happen
We develop the methods for your data, analyze it, and generate figures
STEP5 - Results delivery
We deliver a project report to you in the format you prefer
How it works
Our service is designed for scientists who want easy access to just the right kind of bioinformatics expertise.