Metagenomics
A comprehensive and detailed overview of microbiomes of different origins
Our metagenomics pipeline, is designed to work with both data obtained using shotgun protocols and with selective amplifications (sequencing of 16S, sequencing of 18S, etc.).
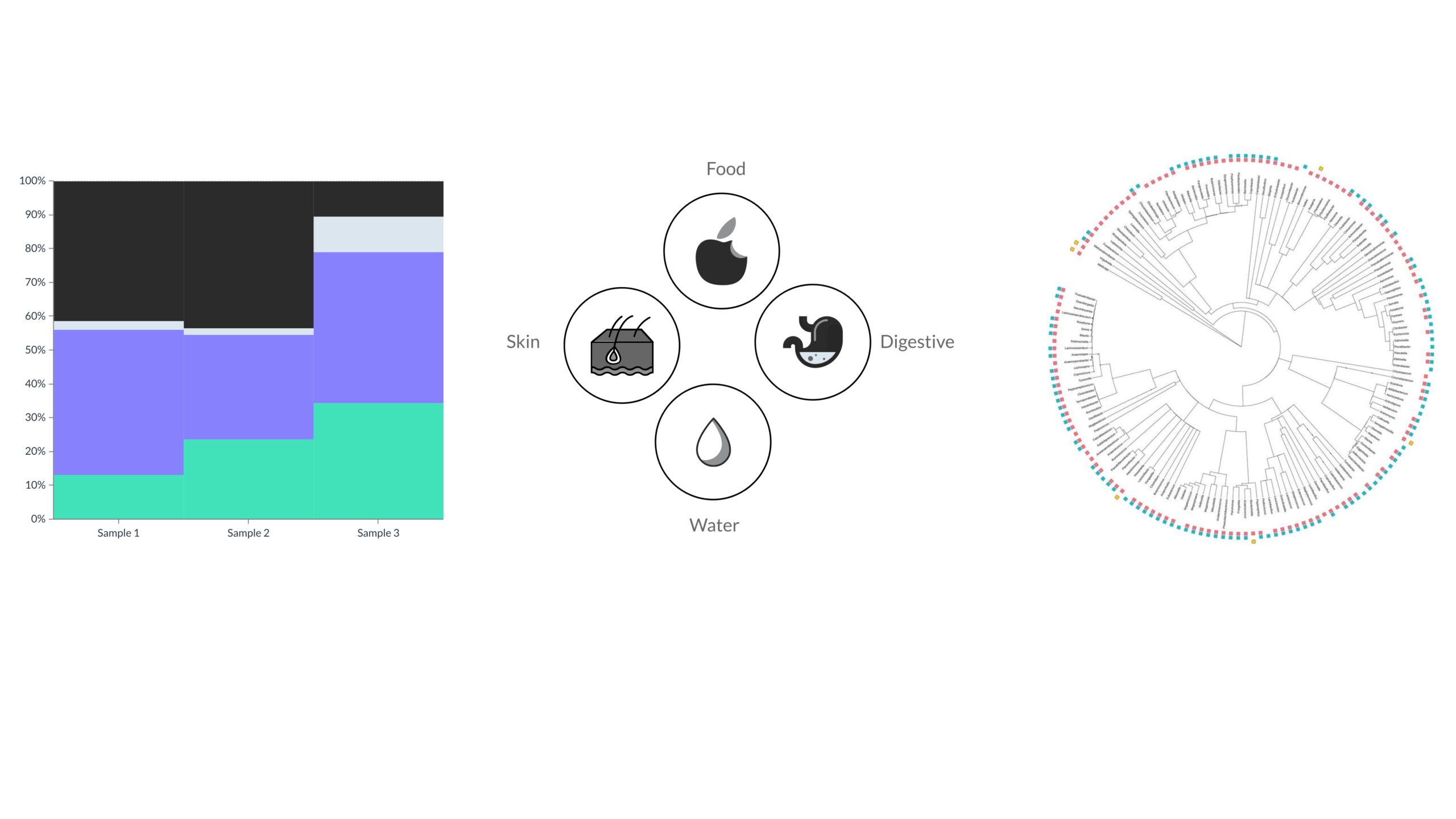
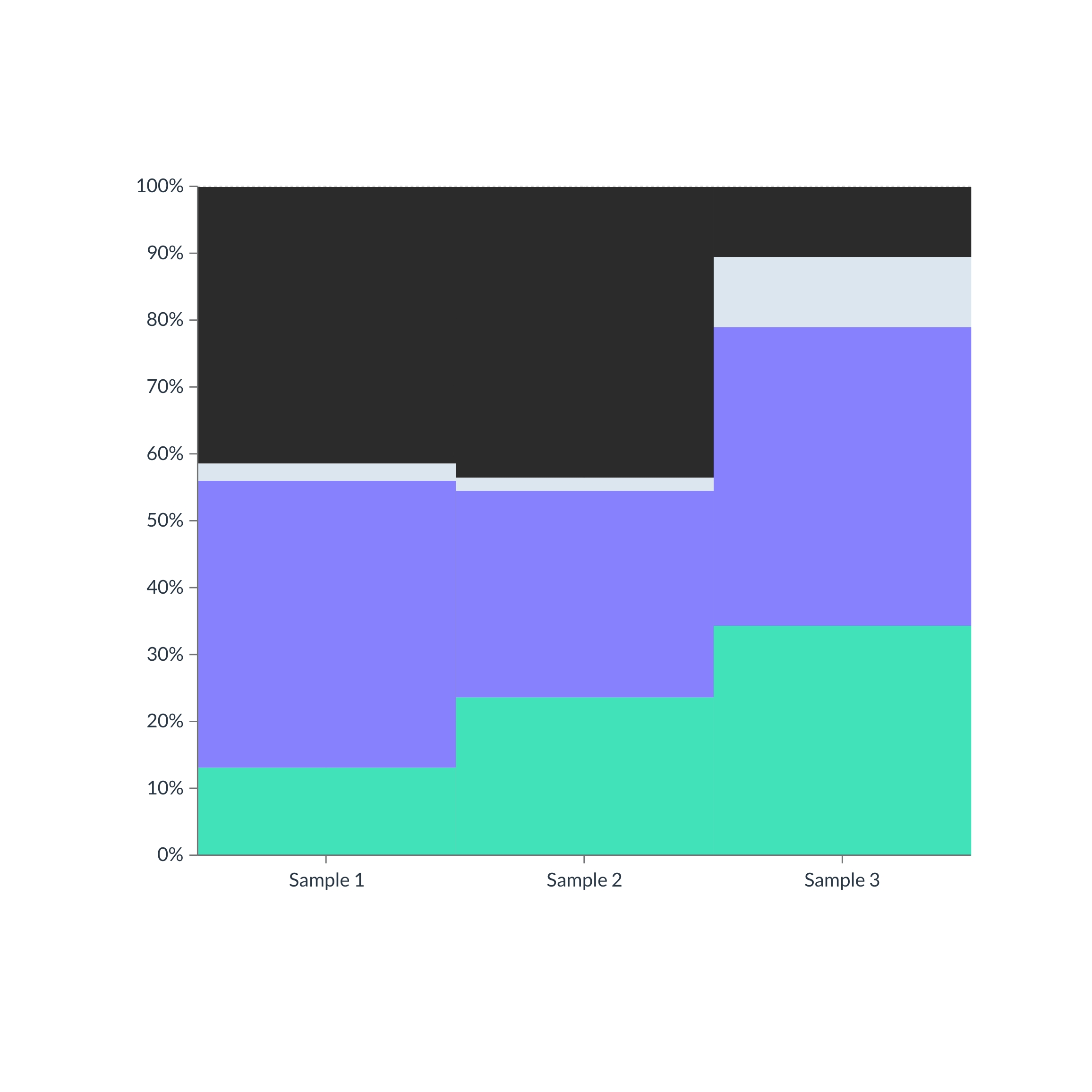
Our pipeline enables us to obtain a comprehensive and detailed overview of microbiomes of different origins: human health (stomach, skin, etc.), agriculture, and the environment (land, water, organic waste, etc.). The scientific classification to work at higher levels (prokaryotes, eukaryotes, and viruses) is integrated with a functional classification.
Metagenomics expertise and consulting
Microbiome research generates a large mass of data by 16S rRNA and shotgun metagenomic sequencing. Efficient bioinformatics pipelines aim to process quickly and accurately sequencing files, however they do not reduce the complexity of the data. Many statistical methods have been developed specifically for microbiome research. Although useful, they are often misused or misunderstood. The results can be felt obscure or untrustworthy, but a specific expertise can clear the way.
We allow our customers to benefit from an experience acquired by analysing dozens of metagenomics datasets generated by 16S rRNA and shotgun sequencing.
16S sequencing
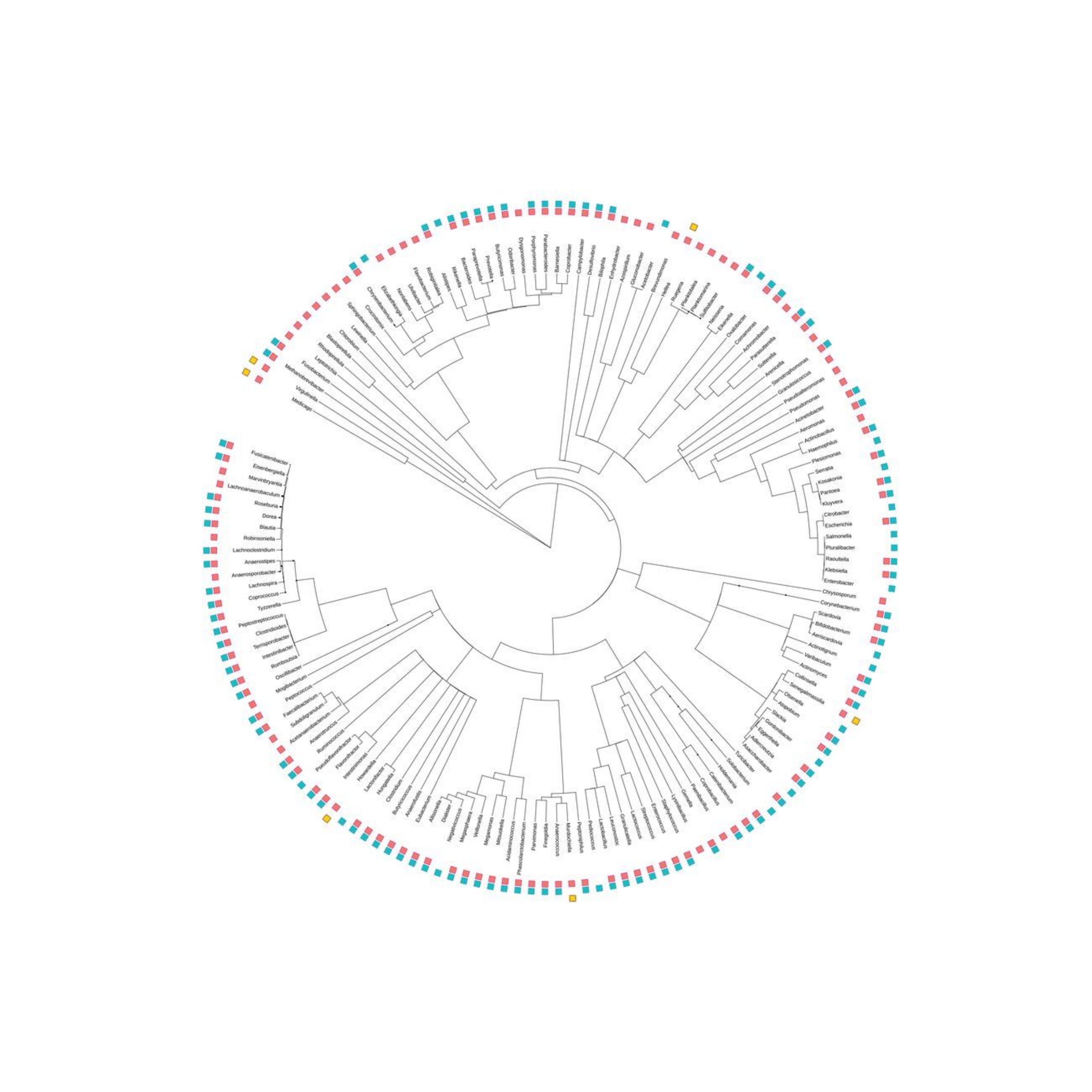
Metagenomic analysis of microbial populations is often performed using the prokaryotic 16s ribosomal RNA (rRNA) gene. These genes contain conserved and variable regions that are studied for phylogenetic classification. In clinical microbiology, molecular identification based on 16s rDNA sequencing is applied fundamentally to bacteria whose identification by means of other types of techniques turns out impossible, difficult, or requires a lot of time.
16s rRNA importance and applications :
-
- 16s rRNA sequencing has been established for the identification and taxonomic classification of bacterial species.
- It can prove to be a method for the recognition of novel pathogens.
- Being the conserved gene in bacterial species it provides the identification of useful signature sequences and patterns.
- 16s rRNA sequencing can lead to the identification of completely new species.
- The conserved variable regions facilitate sequencing and phylogenetic classification.
- Researchers can achieve species level sensitivity for metagenomic surveys of bacterial populations.
- In clinical microbiology, molecular identification based on 16s rDNA sequencing is applied fundamentally to bacteria whose identification by means of other types of techniques turns out difficult, or requires a lot of time.
Shotgun sequencing
Use Shotgun Metagenomic to comprehensively sample all genes of all organisms present in a given complex sample, assess bacterial diversity and detect the abundance of microbes in various environments. Shotgun metagenomics also provides a way to study non-culturablemicroorganisms that are otherwise difficult or impossible to analyse.
Because analysis of metagenomic sequences is complicated due to the complex structure of the data, our Shotgun metagenomic sequencing experts can use new tools and data resources that have been developed to circumvent these complexities to determine which microbes are present in the community. and what they might be do. Additionally, they can apply analytical strategies and tools specific to metagenomic data and the considerations and caveats associated with their use.
Shotgun metagenomic sequencing is a powerful environmental sequencing approach that provides insight into biodiversity and community function.
STEP1 - Contact us
We have a meeting, phone call or visioconference to discuss your project
STEP2 - Estimate and timeline
Our team compiles a project proposal for your project
STEP3 - Accept or reject
The proposal is up for your consideration
STEP4 - Let Magic Happen
We develop the methods for your data, analyze it, and generate figures
STEP5 - Results delivery
We deliver a project report to you in the format you prefer
How it works
Our service is designed for scientists who want easy access to just the right kind of bioinformatics expertise.